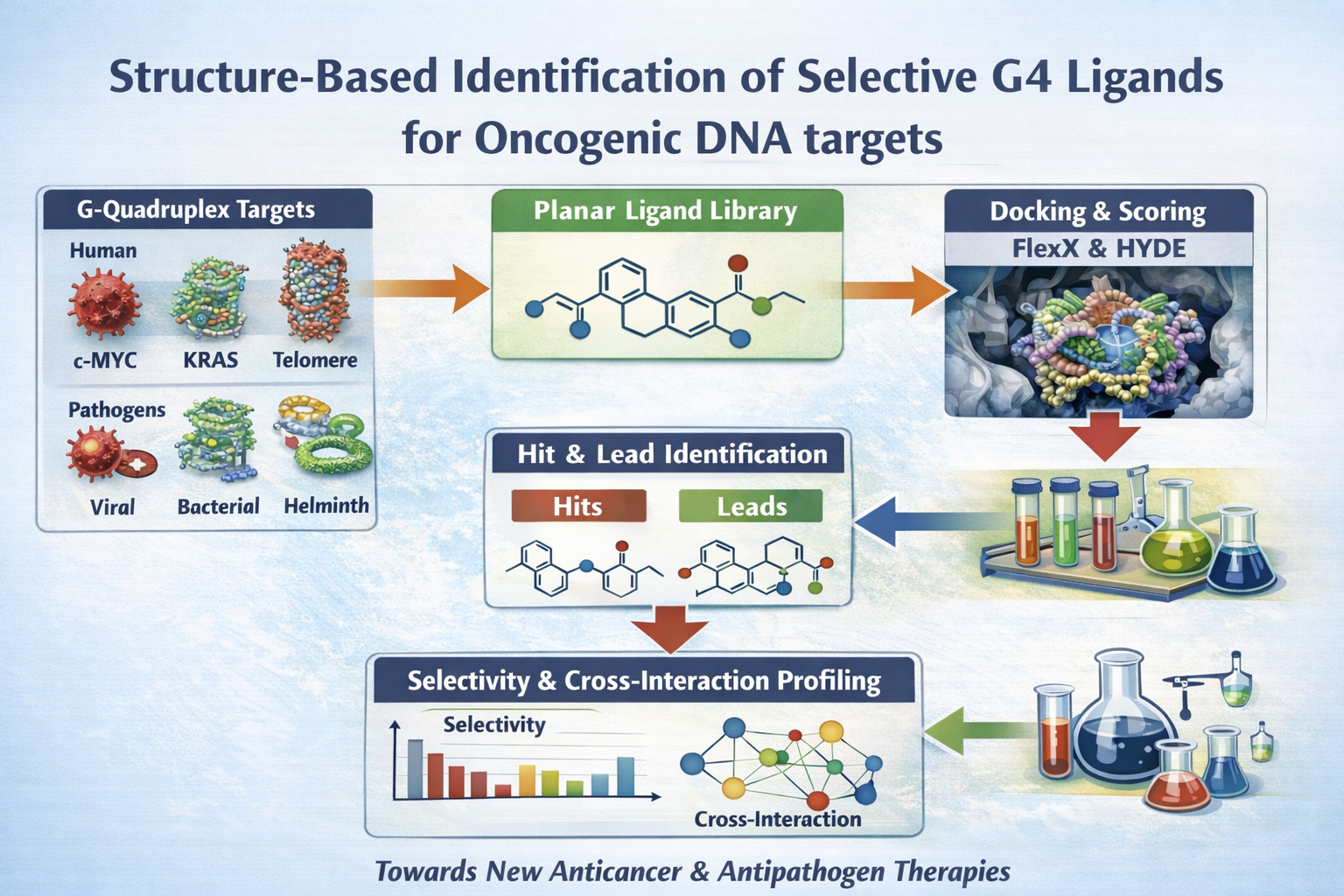
G-quadruplexes (G4s) are non-canonical DNA secondary structures enriched in oncogene promoters and telomeric regions, where they play key roles in transcriptional regulation and cancer progression. However, the rational identification of selective G4 ligands remains challenging due to structural diversity and limited understanding of binding determinants.
In this project, we propose a structure-based docking study of a novel class of planar G4 ligands combining π–π stacking capability with heteroatoms designed to probe hydrogen-bond interactions. Docking and scoring will be performed using BioSolveIT software against a curated panel of G4 targets, including the c-MYC, c-KIT and KRAS promoter G4s. Comparative analysis of binding poses, affinities and interaction patterns will enable the identification of hit and lead compounds and the elucidation of structure–selectivity relationships. Selected ligands will also be evaluated for potential cross-interactions with additional targets.
Alessio Maria intends to achieve the following milestones:
- Selection and preparation of four flagship G-quadruplex targets; setup of the ligand library and definition of docking workflows.
- Docking, pose analysis, and HYDE-based scoring across all targets; comparative selectivity profiling and identification of hit compounds.
- Lead prioritization based on binding poses and selectivity; cross-interaction assessment and generation of hypotheses for chemical synthesis