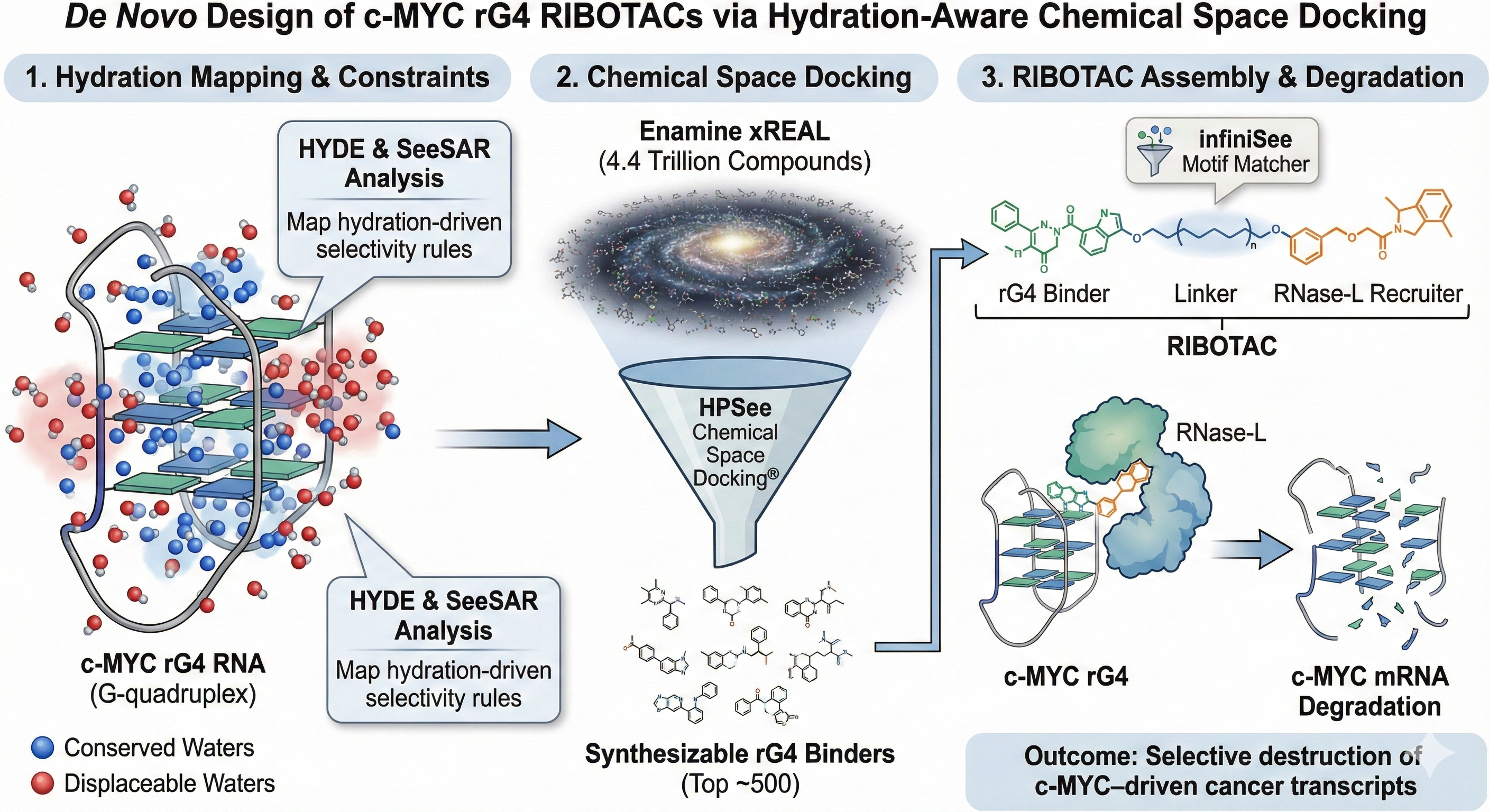
The “RIBOTAC-Space” project aims to degrade c-MYC mRNA by targeting its validated RNA G-quadruplex (rG4) with small-molecule RIBOTACs. RNA binding is strongly influenced by structured water and ion networks, which conventional docking fails to capture. We will use SeeSAR/HYDE to map displaceable vs. conserved waters around the c-MYC rG4 grooves, establishing hydration-driven selectivity rules. Guided by these constraints, HPSee Chemical Space Docking will mine the 4.4-trillion Enamine xREAL space for synthesizable rG4 binders optimized for water-thermodynamic fit. infiniSee’s Motif Matcher will subsequently identify ideal linkers to couple these binders to RNase-L recruiters, enabling selective, proximity-induced RNA degradation. By combining hydration thermodynamics with ultra-large chemical space exploration, this project showcases BioSolveIT’s capability to unlock RNA-targeted therapeutics and advance a high-impact strategy against c-MYC–driven cancers.
Soetsu intends to achieve the following milestones:
- HYDE-based hydration mapping of c-MYC rG4; define water-displacement and groove-binding constraints for selective RNA targeting.
- Run HPSee Chemical Space Docking on Enamine xREAL; select top ~500 synthesizable scaffolds satisfying M1 hydration rules.
- Use infiniSee Motif Matcher to design optimal linkers coupling rG4 binders to RNase-L recruiters for RIBOTAC construction.