In colorectal cancer, elevated levels of 14-3-3ζ are found to enhance cancer cell survival by inhibiting apoptosis. 14-3-3ζ sequesters BAD, a pro-apoptotic member of the BCL-2 family, in the cytoplasm, thereby promoting cancer cell survival. While this represents a key survival mechanism, the potential of targeting 14-3-3ζ:BAD interactions as a chemotherapeutic strategy has remained largely unexplored. In the present work, we conducted an integrated computer-assisted drug discovery process, based on large library shape screening; ligand-based and structure-based pharmacophores; docking, MD simulations, and free energy calculations to find drugs that can selectively disrupt the 14-3-3ζ:BAD complex. This study lays the groundwork for the first detailed CADD study to find therapeutics to break the 14-3-3ζ:BAD duo against colorectal cancer.
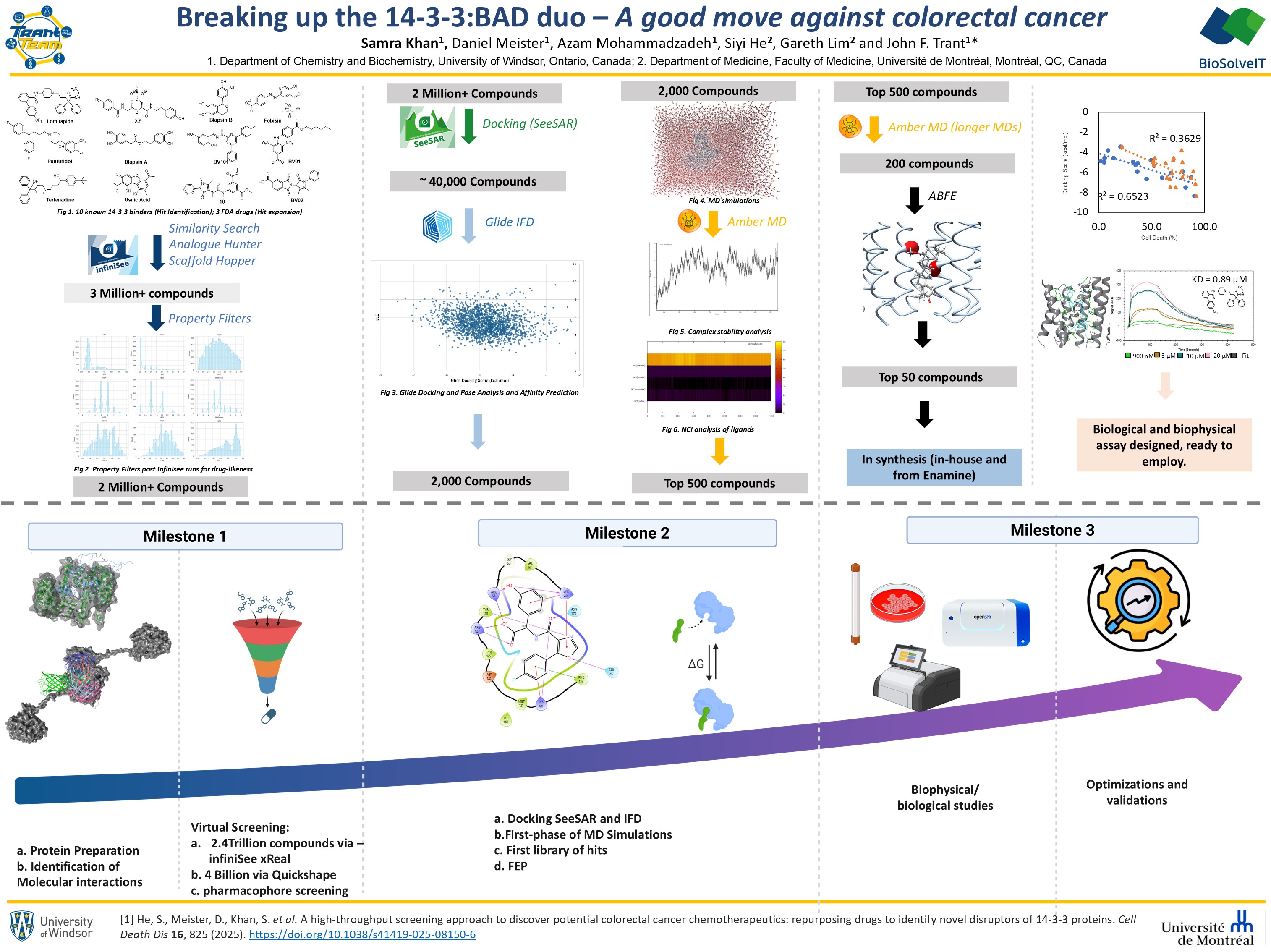
We had two streams 1) we took 10 ligand templates from the literature and conducted a complete Hit identification pipeline and 2) we found 3 FDA-approved drugs in our cell-based screening that induced apoptosis by inhibiting 14-3-3ζ:BAD, and we used those to conduct Hit optimization to further strengthen our Hit pool.
This work would not have been possible without BioSolveIT’s infiniSee xREAL, which provided us with the most efficient ultra-large chemical screening. BioSolveIT’s SeeSAR docked ~2 million compounds fast, while Hyde function allowed us to rank the compounds for the next phases much seamlessly.
Currently, we have completed our computational pipeline for Hit identification, established biological and biophysical assays, and have on-going final phase of our third milestone, which involves synthesizing some top hits in-house, and commercially obtaining others and present workflow and evaluating them using experimentally via our designed assays. In future, we will employ the CSD (chemical space docking) as comparative tool to our current workflow based on brute docking.
Lack of data on compounds with high binding affinity for 14-3-3ζ was a big challenge; and we solved many problems there and established a reference for 14-3-3ζ inhibitor with clear biophysical data. Overall, the project advanced well, usually for this kind of workflow, our estimated timeline is 1+ years for computational work alone, however, BioSolveIT resources reduced our computational timeline to half, and allowed us to focus on experimental aspects.
Lastly, our current results strongly indicate that the crosstalk of our computational and experimental workflows will enable the identification of selective and potent inhibitors capable of disrupting the 14-3-3ζ:BAD duo – representing a ‘good’ move against colorectal cancer therapeutics.
After 1 year, Samra has achieved the following goals:
- Milestone 1 was successfully completed. We used 10 known inhibitors of 14-3-3ζ as query on infinisee xREAL to build libraries of new potential Hits. The vast chemical library in xREAL simplified the compound identification. We used scaffold hopper to identify similar compounds and analog hunter for diverse analogs, generating 200,000 compounds for each, totalling ~ 2 million. We aimed to generate two sets per template: one with high structural similarity and one more diverse (lower similarity) but retaining key chemical features for binding. These were filtered using Optibrium properties to ensure drug likeness. We also used Quickshape on a prepared library, but it runs slower and used a 4 billion compound subset of REAL; hence, compound similarity was lower than with xREAL. Afterwards, we prepared the 14-3-3ζ protein model (PDB 2C1J, chosen for no missing parts), identified the binding pocket, and got it ready for milestone 2.
- Milestone 2 was also successfully completed. First, we performed docking on the ~2 million compounds from Milestone 1 using SeeSAR. Binding modes and affinities were compared with known inhibitors and filtered based on predicted binding, chemical properties (e.g., clogS and LogP), and poses (e.g., interactions and torsion quality), resulting in several thousand compounds with significantly improved predicted binding. Overall, binding affinities improved from high µM for templates to pM–low nM for top hits. We also performed induced-fit docking with IFD in Glide for further refinement; due to long runtimes, only selected top compounds were being docked (~40,000 compounds). MD simulations were run on top compounds from this milestone (~2,000 compounds at 50 ns), which were scored using MM-GBSA. The top 500 compounds were advanced to the next phase.
- We successfully completed 1st phase of this milestone: In the 1st phase, 500 compounds from Milestone 2 were subjected to longer MD simulations, top 200 were taken to calculate ABFE values using TI, and the top 50 candidates were ranked. In the 2nd phase, we developed cell-based assays to evaluate 14-3-3ζ inhibition in CRC cells, and tested top-ranking compounds using SPR, resulting in validated protocols and hit identification of three FDA-approved drugs: Lomitapide, Terfenidine & Penfluridol. The 3rd phase is ongoing, the three drugs were used as templates for hit expansion using infiniSee xREAL7, yielding ~1M compounds, which we are currently running through the workflow. We are also currently synthesizing 50 hits, some in-house and some through Enamine. Completion of this milestone will link our computational predictions with experimental validation; helping us complete a story of integrated workflow on finding selective and better disruptors for 14-3-3ζ:BAD complex.