In the first three months of the project, we have rapidly progressed toward the development of an optimal target structure for docking and the initiation of the AI-accelerated virtual screening campaign. We have prepared the Enamine REAL library for Deep Docking, and we have characterized the binding of acridine compounds into the XPF binding pocket. A major challenge we fronted was the low performance of docking tools to correctly place and rank the acridine derivatives, which made difficult to obtain clear structure-activity relationship. However, by using more than one XPF structure, we were able to develop a better scoring strategy, which also differentiated decoys from actives in docking scenarios. We are now well-positioned to initiate the screening campaign. We also established a collaboration with experimental groups that will test our prioritized compounds in enzymatic assays as well as in combination with cisplatin in cancer cells.
After 3 months, Francesco has achieved the following milestones:
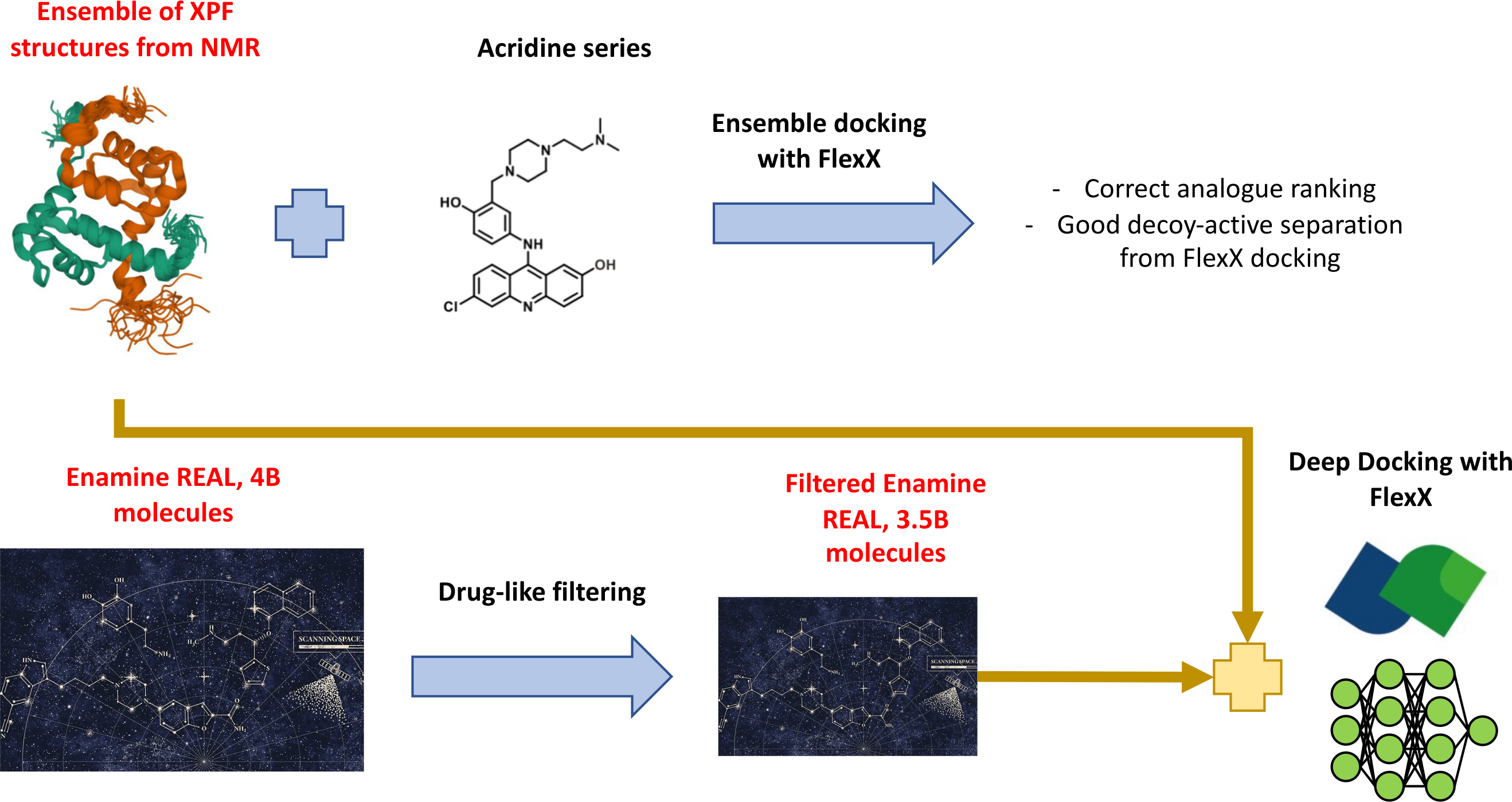
- Using a single XPF structure to rank different acridine analogues on the basis of their predicted affinities led to large disagreement with experimental data. We have taken the approach of using multiple XPF structures in an ensemble docking scenario, by removing redundant conformations from the NMR structural ensemble, and we observed that this led to better results both in terms of analogue ranking and in comparison with decoy screening, where decoys were generated with DUD-E and docking was performed with FlexX.
- See above. We are currently exploring other PPI sites similar to the XPF pocket to further extend our decoy-active screening exercise. The use of Deep Docking will tremendously facilitate the adoption of ensemble docking-based screening of the REAL database, which would be otherwise intractable.
- We have prepared the library by applying a series of preliminary drug-like filters to remove compounds with high number of chiral centers, low carbon sp3 fraction, many rotatable bonds, and so on. We have calculated dominant tautomer/ionic forms for each molecule in the library, and calculated their circular fingerprints (1024 bits, radius 2). We are currently calculating other chemical fingerprints to compare Deep Docking performances and adopt the most efficient screening strategy for FlexX docking.