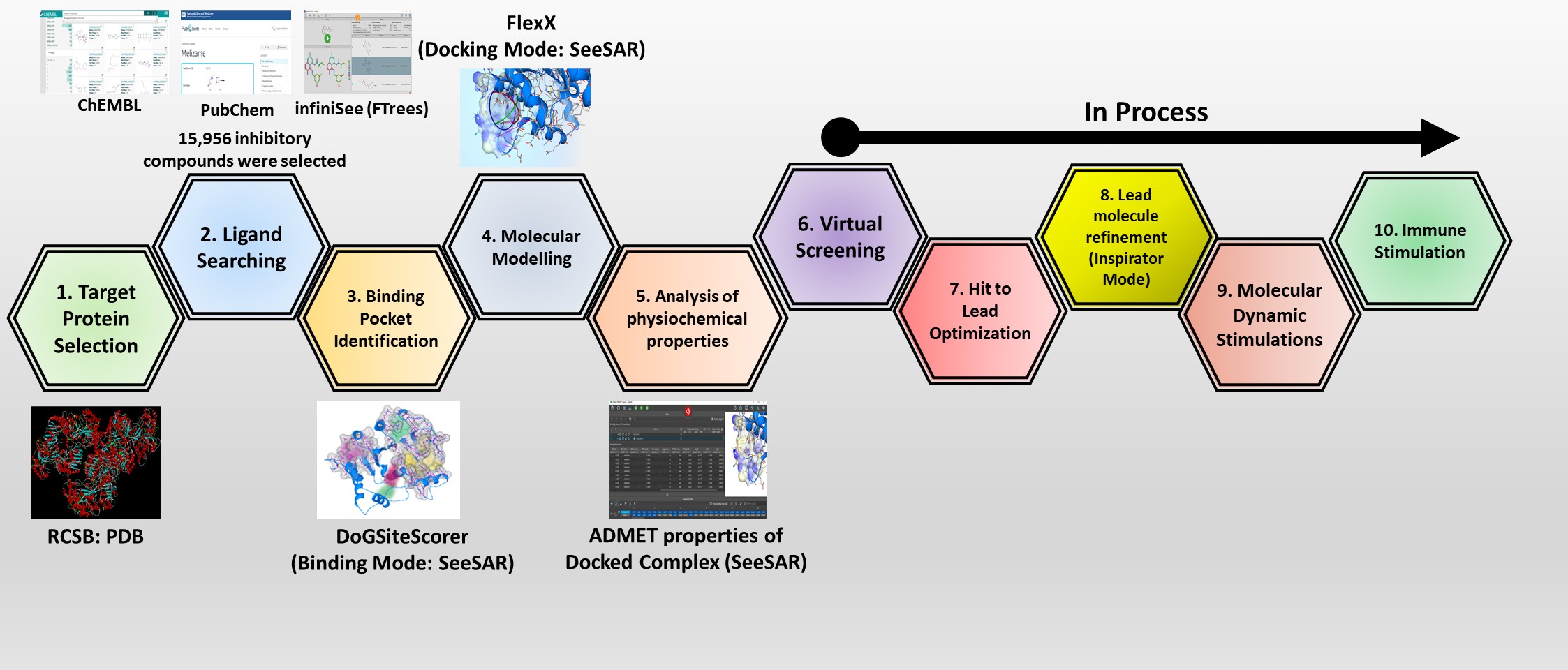
The prime objective of the present research is the recognition and synthesis of suitable lead molecules for the inhibition of non-structural protein (NSP) of zika virus (Yellow fever) that will ultimately help them in the prevention of virus replication. In the first three months of this research project, our aim was to screen appropriate inhibitors against NSP of zika virus. Initially, the selection of both biologically active and inactive compounds was done from literature and ligand screening databases like CHEMBL and PubChem. The selected inhibitory compounds were further screened on the basis of their 2-D structure similarity search by creating real chemr 2-D structure similarity search by creating real chemical spaces using FTrees (infiniSee). The target protein structure was obtained from RCSB Protein Data Bank and analyzed for both occupied and unoccupied binding site pockets using DogSiteScorer (SeeSAR). Furthermore, molecular modeling interaction studies were also analyzed.
After 3 months, Muhammad has achieved the following milestones:
- In the first month of our study, the biologically active and inactive inhibitors were obtained from literature and various online databases according to our selected viral target protein which was retrieved from RCSB Protein Data Bank. The structures of the selected ligands were downloaded in SDF (spatial data file) format using PubChem (https://pubchem.ncbi.nlm.nih.gov/), ChEMBL (https://www.ebi.ac.uk/chembl/). BioSolveIT’s entirely user-friendly and efficient tools SeeSAR and infiniSee were successfully used for the ligand and protein preparation. The target protein was analyzed in protein mode by loading and superposing the protein structure. In the binding site mode, the target viral protein was analyzed for the binding site pockets. Under the docking mode, the selected inhibitors were docked with the target viral protein, thus forming the various poses. After standard molecular docking, the ligands were screened on the basis of best pose formation with the protein binding sites.
- During the second month, the final ligand library was formed on the basis of best binding site interactions of inhibitory molecules with the target viral protein. The selected best pose along with estimated binding affinities of inhibitor with viral target protein was analyzed for the development of pharmacophore. The ligands with the best poses and pre-eminent binding affinities were selected which were further analyzed for physicochemical properties thus leading to the generation of final ligand preparation. FTrees (infiniSee) was explored to analyze chemical spaces, 2D similarity and active parts of the compounds, and libraries were generated.
- In the third month of this research project, we are analyzing the best lead molecules by improving the molecular interactions of the inhibitors with the target viral protein using molecular editor mode. The editor mode of SeeSAR enables us to eliminate clashes by removing or adding the atoms, improves the binding affinity and torsion angles between the final lead molecules with viral protein binding sites. After removing the clashes inspirator mode of SeeSAR is used for further refinement of the lead molecules. The inspirator mode will be used for the elimination of the unfavorable atoms in the docked conformers with the target binding protein. In the next nine months, we will work on lead optimization using molecular editor and inspirator mode of SeeSAR. The molecular dynamic simulations will be performed to analyze the various conformational changes of the docked complex and immune stimulations to analyze the computational immune response generation against the best lead molecule.