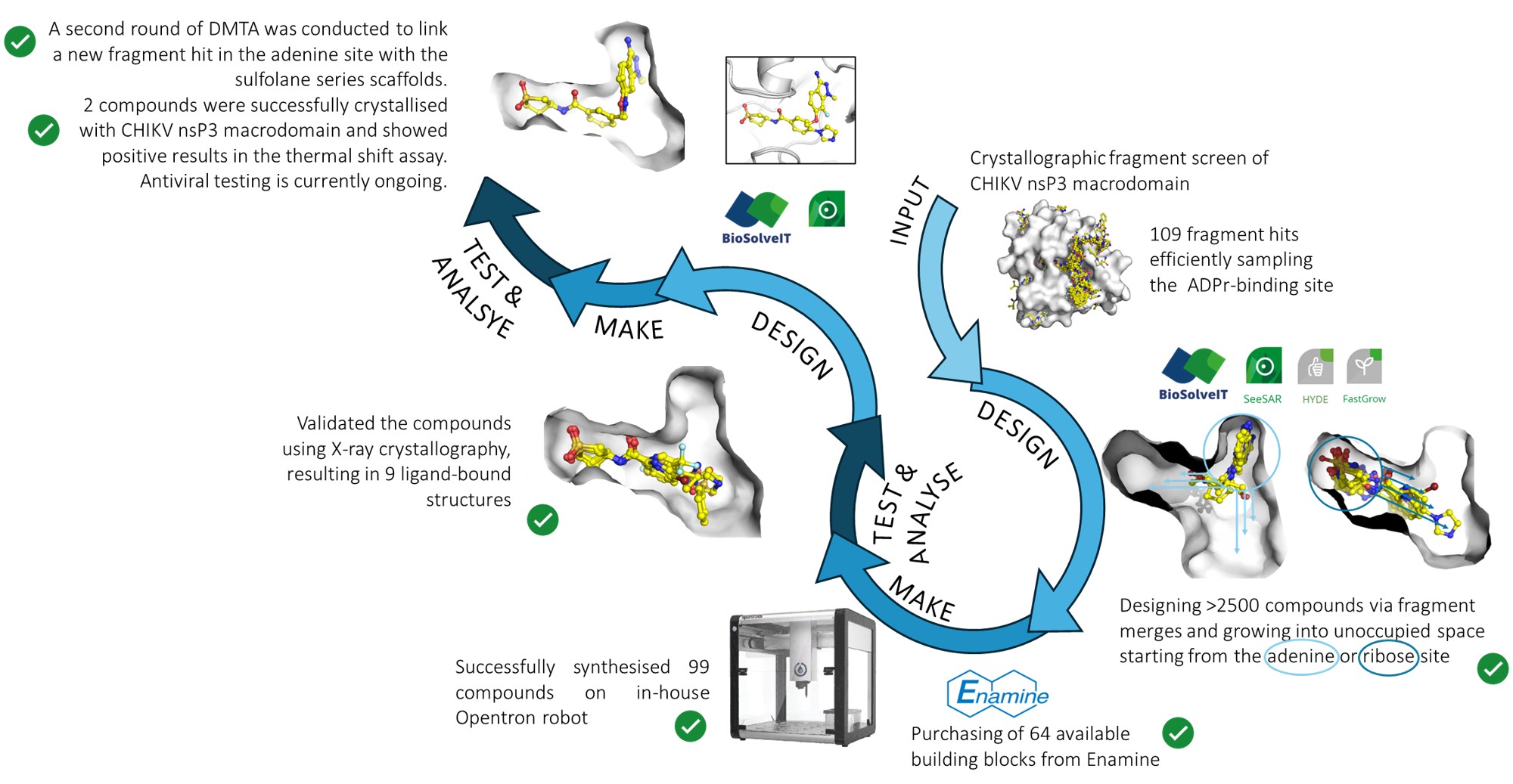
Using BioSolveIT, we designed compounds based on the previously obtained fragment screen hits or early validated algorithmic fragment merges that could be easily synthesised via amidation reactions on our in-house Opentron robot. We generated the compounds via fragment merging, linking or growing starting from two different sites: from the adenine site towards the oxyanion site or phosphate site (‘pyrrolopyrimidine series’); and from the ribose site towards the phosphate site and oxyanion site (‘sulfolane series’). This resulted in 1012 and 1548 potential scaffolds for the sulfolane and the pyrrolopyrimidine series, respectively, which would be generated from 438 building blocks (acids and amines). Unfortunately, due to availability, we were only able to purchase 64 of the building blocks, which were used to synthesise ~100 pyrrolopyrimidine and ~150 sulfolane series compounds.
In the end, we successfully synthesised 74 pyrrolopyrimidine and 25 sulfolane series compounds according to LC-MS analysis. The crude reaction mixtures of the compounds underwent soaking into protein crystals and X-ray crystallographic analysis showed successful binding of 9 compounds of the sulfolane series. Unfortunately, none of the 'pyrrolopyrimidine' compounds bound, but some of the building blocks bound to the adenine site, as would be expected.
We additionally performed a buffer screen to investigate the stability of the protein in different buffer conditions to find a suitable condition to be used in the planned biophysical assay (GCI analysis). This revealed that a higher salt concentration (300 mM vs 100 mM NaCl) increases the stability of the protein.
In the meantime, collaborators at UCL performed an NMR fragment screen and found an additional fragment hit binding to the adenine site, which was also verified using X-ray crystallography. We therefore designed some additional compounds that linked our sulfolane series scaffold with this hit. These compounds were synthesised at UCL. We obtained protein-ligand structures, and the best compound displayed an increase of 4°C in the thermal shift assay.
These compounds are currently undergoing antiviral testing, while we are still working on generating a successful GCI method in the meantime to analyse all synthesised compounds.
After 1 year, Jasmin Cara has achieved the following goals:
- We successfully used the BioSolveIT software suite, including SeeSAR, to design over 2500 scaffolds for potential follow-up compounds, based on 438 building blocks (acids and amines). However, due to the low availability of the building blocks, we would have been able to synthesise only about ~250 compounds in total.
- In the end, we successfully synthesised 99 compounds using the OpenTron, which is part of our Chemist-Assisted Robotics (CAR) and which were verified using LC-MS analysis. The compounds underwent soaking and X-ray crystallography to obtain ligand-bound protein structures, which was successful for 9 sulfolane series compounds.
- This information was used to further design 4 follow-up compounds by linking the sulfolane series scaffold with an additional hit obtained in the adenine-binding site of CHIKV nsP3 macrodomain, which were synthesised at UCL. We have obtained ligand-bound structures for two of the four designs, which also correlate with the thermal shift data. The compounds currently undergo antiviral testing. We are still optimising the biophysical assay to generate affinity data for all of the synthesised compounds, and will then publish everything together. In the meantime, we have shared all structures on our interactive web platform Fragalysis: https://fragalysis.diamond.ac.uk/viewer/react/preview/target/CHIKV_Mac/tas/lb32633-6