Download SeeSAR 15 'Apollo' Now
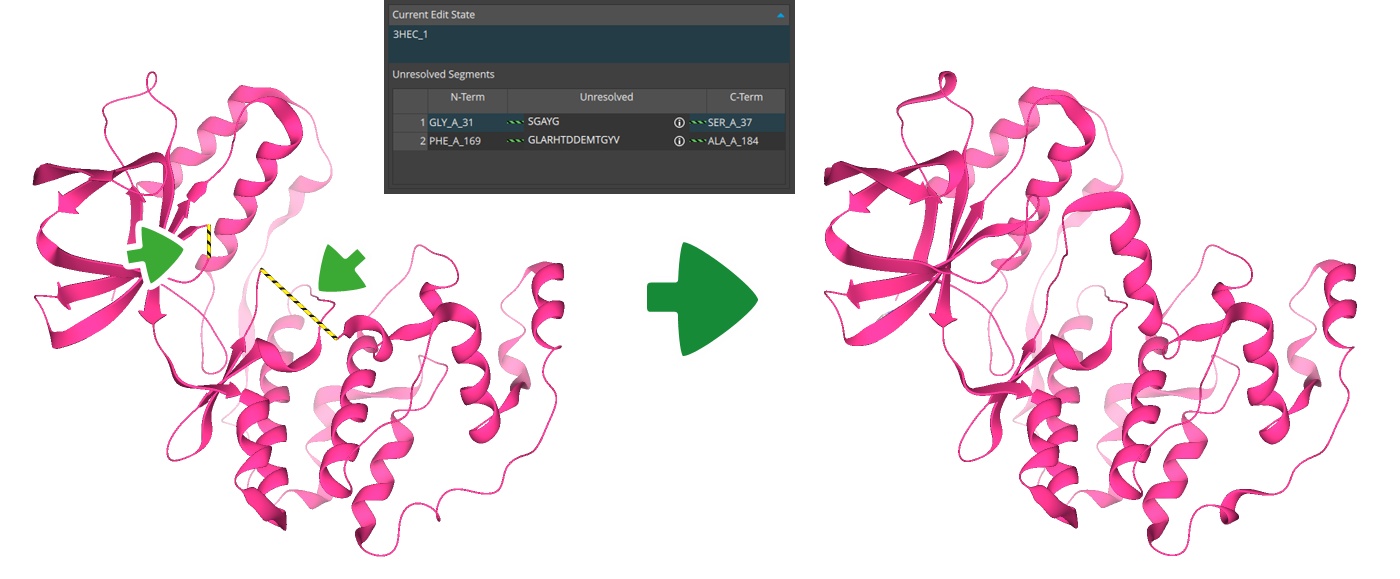
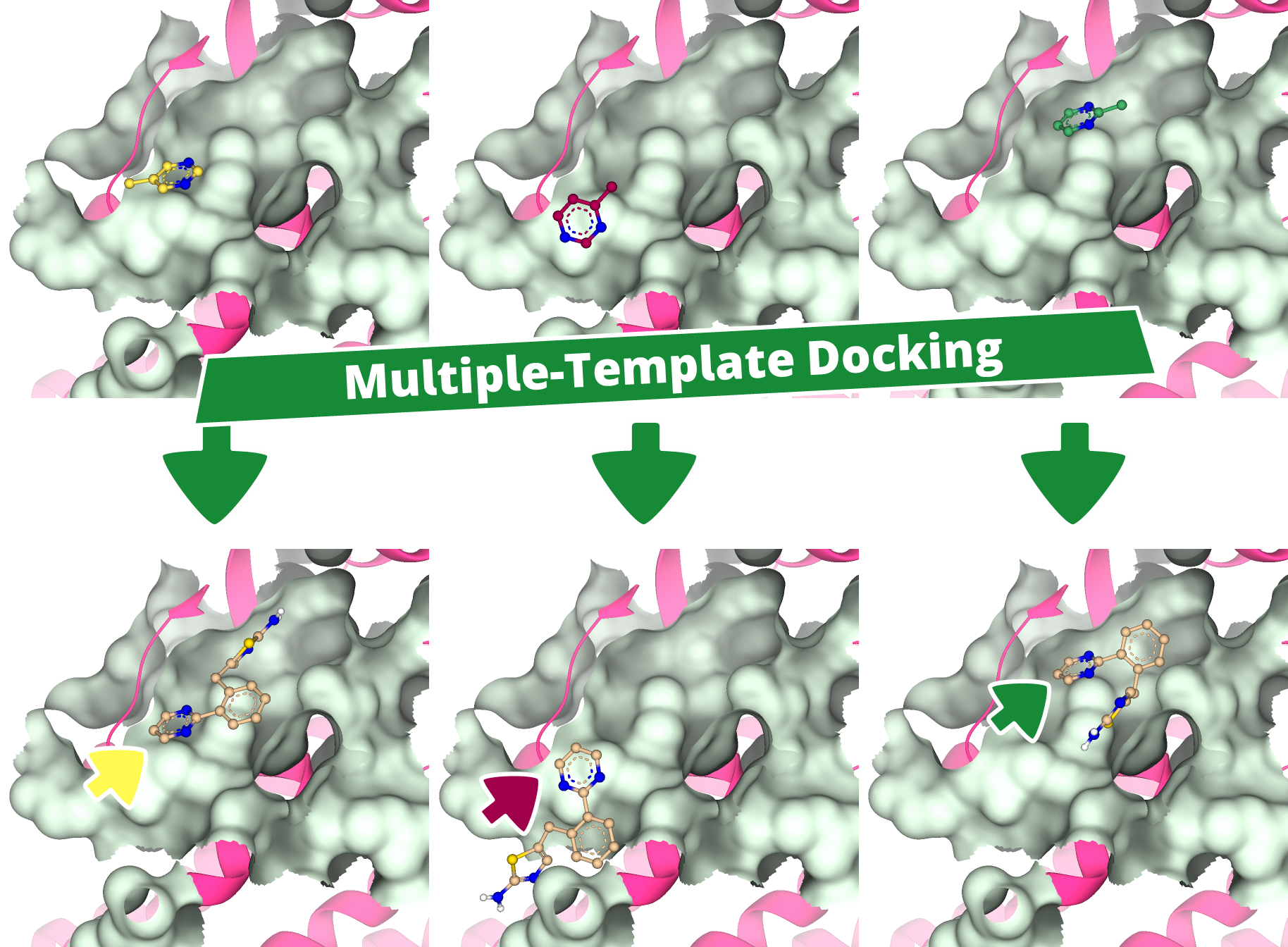
This concludes our overview of the features introduced in the SeeSAR 15 'Apollo' release. A comprehensive changelog, which details all functional additions such as the YASARA-powered loop modeling and multiple-template docking, is available online with the new version.
We hope these highlights have showcased how this major update can advance your drug discovery projects. The full version of SeeSAR 15 is available now to streamline your design and optimization workflows.
The full changelog, including all new additions and quality-of-life improvements, can be accessed at
this link.