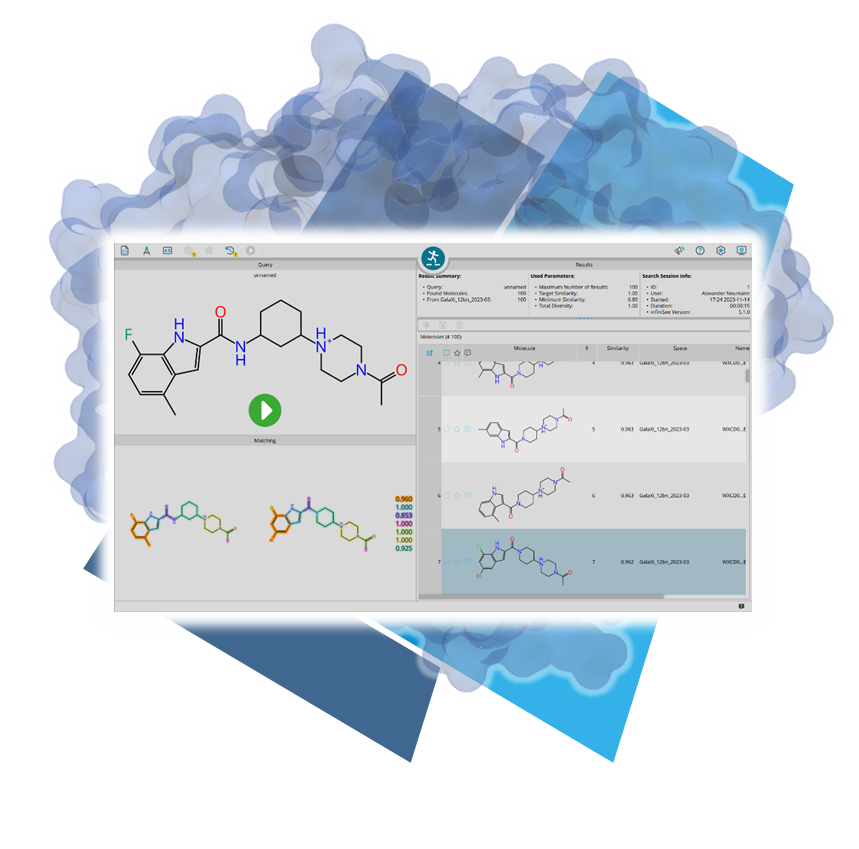
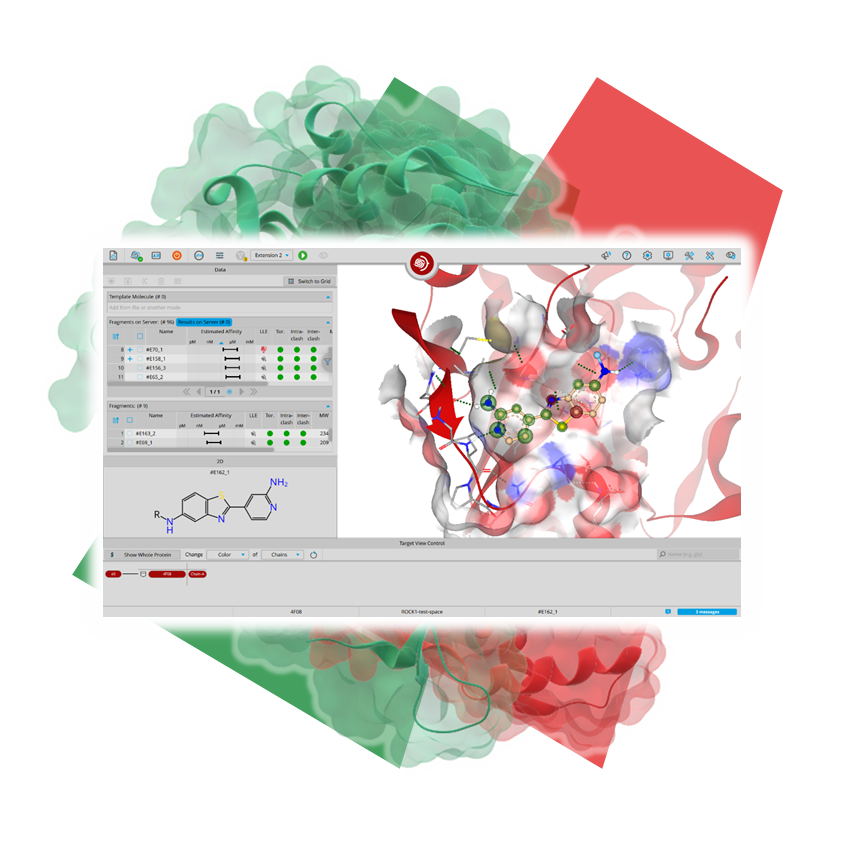
SeeSAR
Drug Design Dashboard

| Version | Building blocks used | Number of products | Products without KILL | Computational time* | Storage size |
| SAVI-Lib-2020 (enumerated library) | 1.55 × 105 | > 1 × 109 | — | HPC (millions CPU h) | ~210 GB |
| SAVI-Space-2020 (Lib-2020 rules) | 1.39 × 105 | 2.34 × 109 | 3.65 × 109 | 3 h | 2.1 GB |
| SAVI-Space-2020 | 1.38 × 105 | 2.40 × 109 | 3.34 × 109 | 3 h | 0.8 GB |
| SAVI-Space-2024 | 2.56 × 105 | 7.55 × 109 | 1.07 × 1010 | 10 h | 1.4 GB |